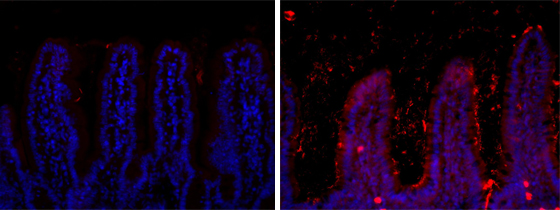
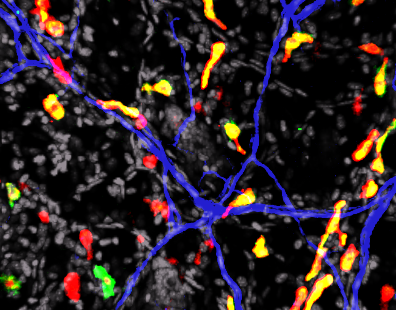
Localisation of the bacteria the ileum of a mouse fed a standard diet (left) or a high-fat diet (right).
© Institut Pasteur
A disruptive element such as a change in diet, and the whole intestinal flora becomes disrupted, with possible repercussions for health. An international research study conducted by the Molecular Microbial Pathogenesis Unit (Institut Pasteur/Inserm), directed by Philippe Sansonetti, has just demonstrated, in mice, the direct influence of a diet too high in fat on the intestinal flora and its environment. In response to this new diet, the bacterial communities reorganise themselves and the small intestine itself is transformed. And this happens in the first month. These results were published in the journal PNAS on 16 September.
The billions of bacteria that populate our intestine – known as the microbiota – play a central role in digestion, but also have a role in some diseases such as type 2 diabetes or obesity. These diseases have often been associated with an imbalance of the intestinal flora, with some bacteria becoming clearly predominant, and a permeable intestine, likely to release inflammatory substances into the bloodstream. But although many studies have been conducted on the state of the microbiota once the disease has become established, few have focused on the development of this intestinal imbalance, e.g. when a high-fat diet is introduced. For this reason, an international research team has addressed the question. “We wanted to see, from an early stage, how the intestinal bacteria behave in the presence of a high-fat diet,” emphasises Thierry Pédron, a research engineer in the Molecular Microbial Pathogenesis Unit (Institut Pasteur/Inserm). “And we soon focused our research on the small intestine, because that was where we saw the most obvious variations!”
Some mice in the study were therefore fed an ordinary diet, whereas others were fed a diet containing 70% lipid. Using genomic techniques, the researchers were able to identify the different bacterial species contained in faecal samples, and monitor the changes in composition of the microbiota in time. They also localised and accurately identified the bacteria within the small intestine.
“Only one month after starting this new high-fat diet, we observed changes in the composition of the microbiota,” says Thierry Pédron. “Some bacterial species proliferated, whereas others declined, and the species Candidatus arthromitus actually disappeared completely. Furthermore, and as never seen before, we observed a massive concentration of bacteria between the intestinal villi.”
Ordinarily, bacteria cannot come close to or even cross the intestinal wall because the epithelium releases antimicrobial peptides and is lined with a protective mucus. The researchers then focused on these intestinal wall defences: they found that production of antimicrobial peptides fell following massive ingestion of fat, and that the mucus layer became thinner. In other words, not only does the microbiota become reorganised under the influence of lipids, but the intestine itself undergoes transformation. And the modifications do not end there. Additional measurements made it possible to demonstrate an increase in the permeability of the small intestine, and a reduction in PPAR-γ activity. “PPAR-γ is a molecule with many functions. It plays an important role in fatty acid metabolism, as well as in inflammation and embryonic development,” explains Thierry Pédron. “This drop seems closely related to the drop in antimicrobial peptide level.” And although the connections between all these results and their potential involvement in some dietary imbalances have not yet been established, it is reassuring to note that when the mice are put back on a balanced diet, everything returns to normal within a month!