Researcher Contact
Brice Felden
Directeur du laboratoire « ARN régulateurs bactériens et médecine »
(U 1230 Inserm/Université de Rennes 1)
E-mail : rf.1senner-vinu@nedlef.ecirb
Téléphone : +33 (0)2 23 23 48 51
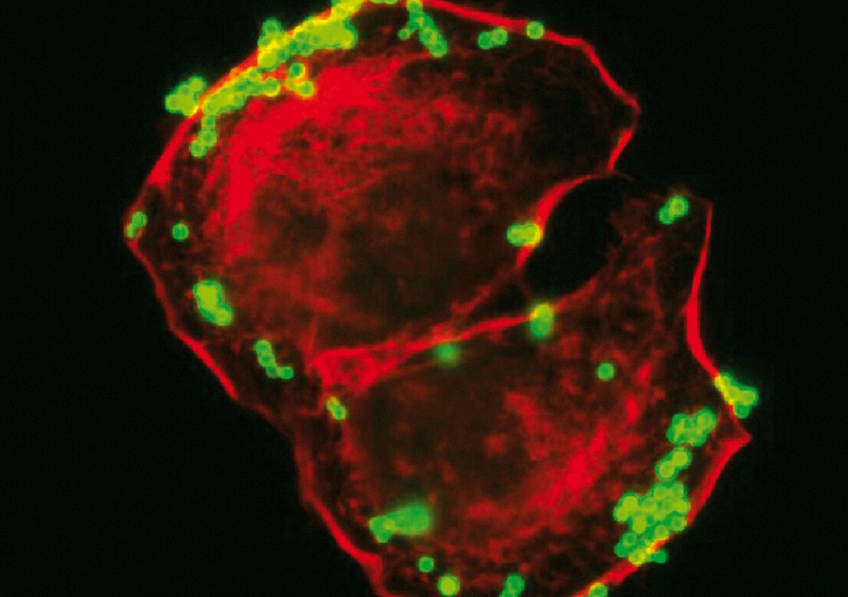
Staphylococcus aureus bacteria (in green) adhering to keratinocytes (in red). © Inserm/Tristan, Anne
In this context, persistence denotes the ability of bacteria to survive high doses of antibiotics without actually becoming resistant. They become persistent by slowing their growth – a bit like going into “hibernation” – to protect themselves from antibiotic treatments. The presence of such antibiotic-tolerant bacteria represents a major public health problem. When the intake of antibiotics is stopped, some of the bacteria “wake up” and are liable to multiply again, presenting a very high risk of relapse or of developing a chronic bacterial infection.
Many of the mechanisms leading to the formation of persistence remain unknown. In their study, the Inserm and Université de Rennes 1 researchers at the Bacterial Regulatory RNAs and Medicine laboratory focused on Staphylococcus aureus. This bacterium is the leading pathogen responsible for nosocomial infections (infections contracted in hospital) and is also implicated in many cases of food poisoning.
In their study, the researchers focused on a Staphylococcus aureus non-coding RNA, i.e. one that is not translated into proteins.
They have shown that once positioned on the ribosomes[1] of the staphylococci, this RNA (referred to as SprF1 antitoxin) reduces protein synthesis during the growth of the bacterium (the aforementioned “hibernation” phenomenon). This mechanism promotes the formation of persistent staphylococci that become insensitive to antibiotics.
“We have revealed an RNA-guided molecular process in which the interaction between this SprF1 RNA and the ribosome is involved in the formation of antibiotic-tolerant bacteria, which are themselves largely implicated in chronic staphylococcal infections,” emphasizes Brice Felden, the professor at Université de Rennes 1 who supervised this work.
These findings also make it possible to envisage a new class of anti-infectives that target persistent bacteria, and thus new treatments for chronic Staphylococcus aureus infections. “Based on these results, we want to develop molecules that fight the persistent bacteria by targeting the SprF1 antitoxin. This strategy is therefore aimed at supplementing the therapeutic arsenal available to clinicians, who find themselves confronted with an increasing number of cases of chronic bacterial disease,” declares Marie-Laure Pinel-Marie who coordinated this research.
These findings have been the subject of a European patent application.
[1] Particles present in all cells that are the “factories” for making proteins.
Brice Felden
Directeur du laboratoire « ARN régulateurs bactériens et médecine »
(U 1230 Inserm/Université de Rennes 1)
E-mail : rf.1senner-vinu@nedlef.ecirb
Téléphone : +33 (0)2 23 23 48 51
RNA antitoxin SprF1 binds ribosomes to attenuate translation and promote
persister cell formation in Staphylococcus aureus.
Marie-Laure Pinel-Marie1*†, Régine Brielle1*, Camille Riffaud1, Noëlla Germain Amiot1, Norbert Polacek2, and Brice Felden1†
1 Inserm, BRM [Bacterial Regulatory RNAs and Medicine] – UMR_S 1230, 35000 Rennes, France.
2 Department of Chemistry and Biochemistry, Bern University, 3012 Bern, Switzerland.
Nature Microbiology, janvier 2021.