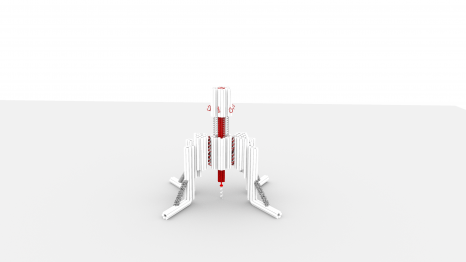
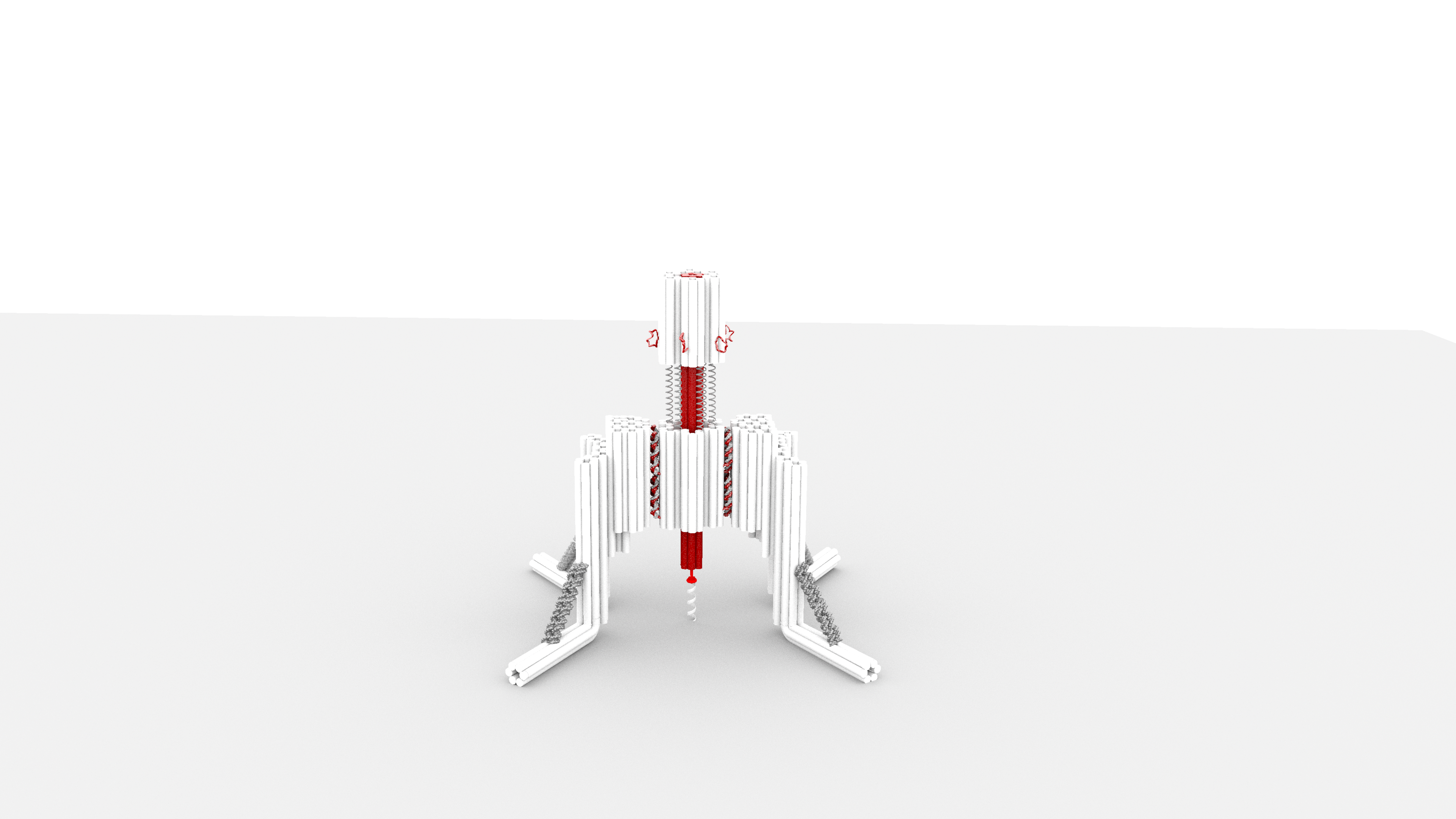
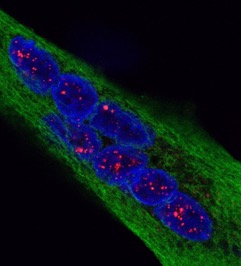
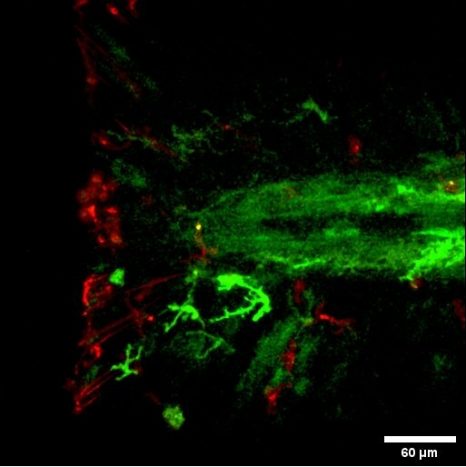
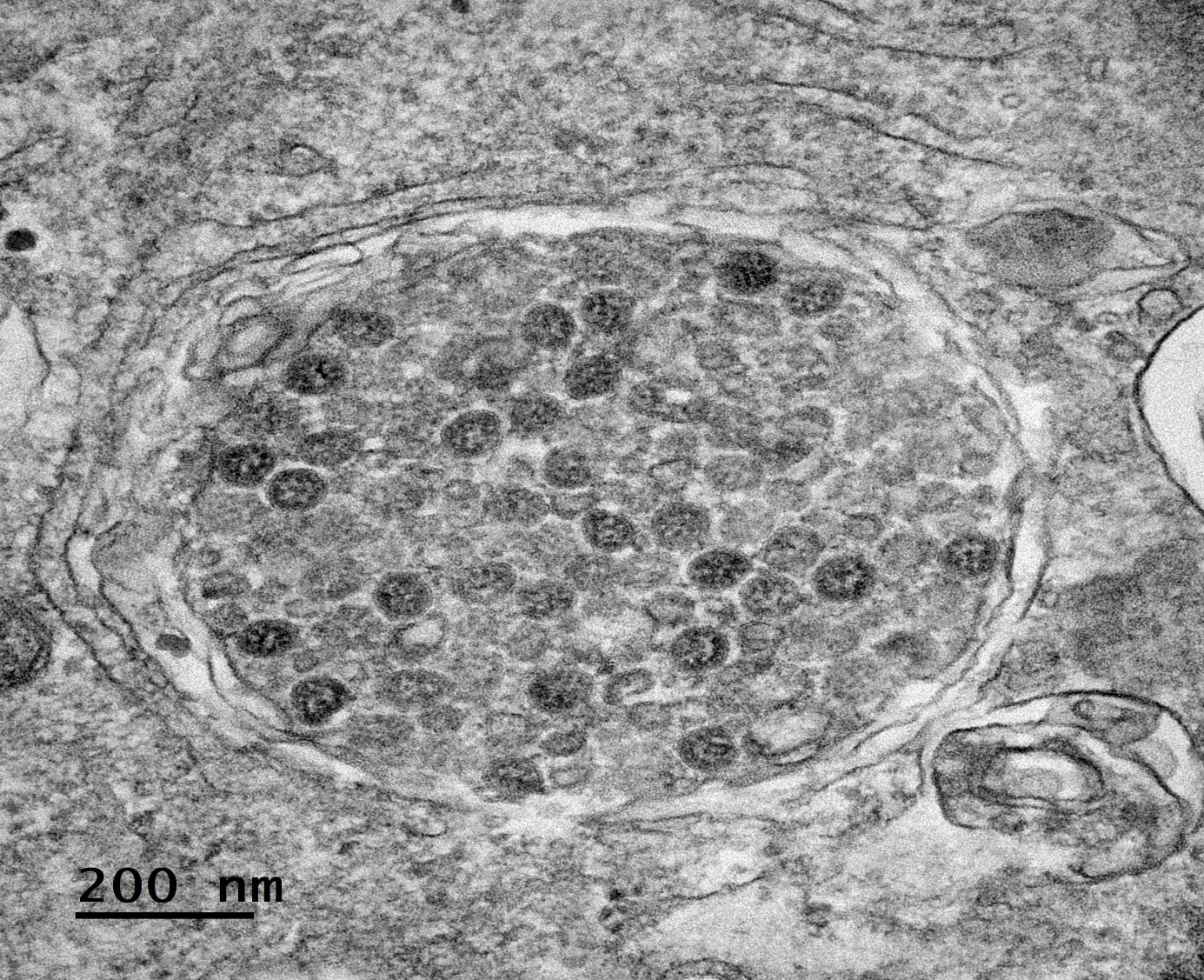
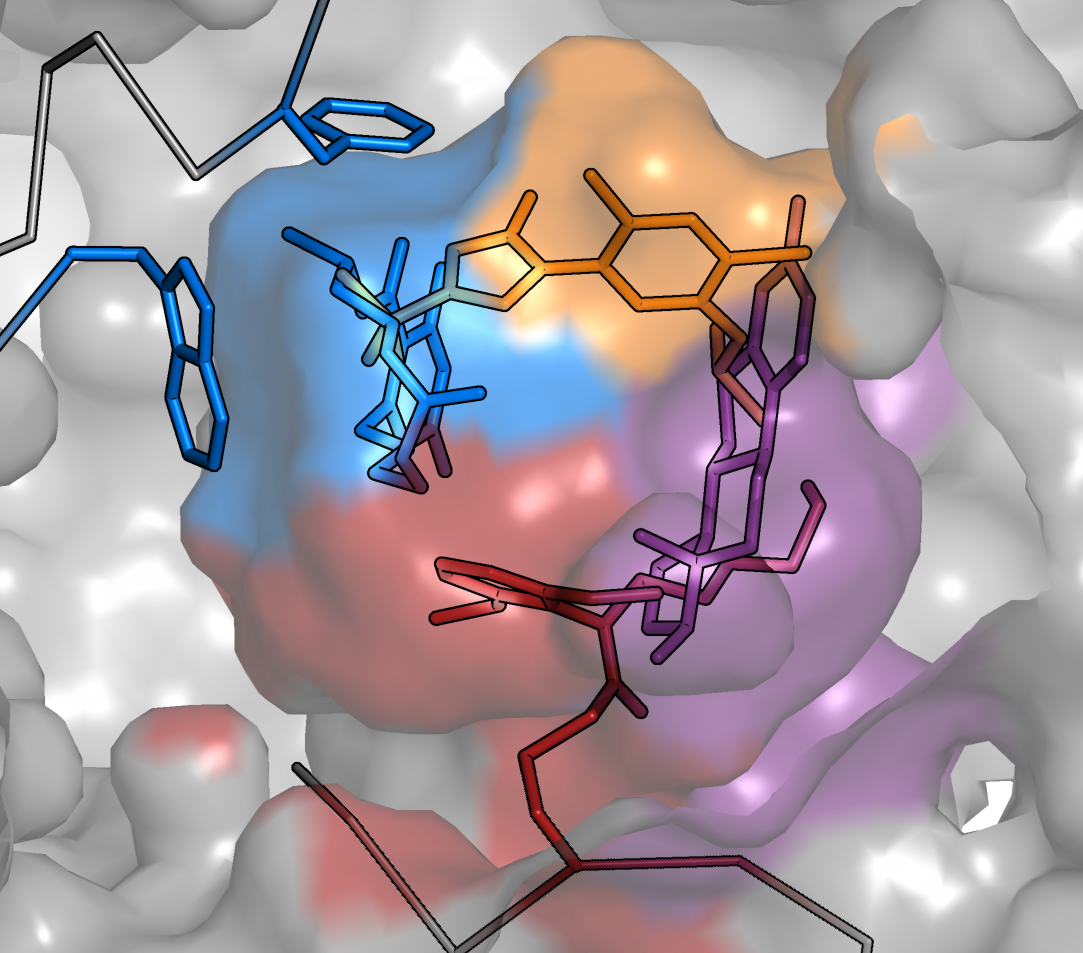
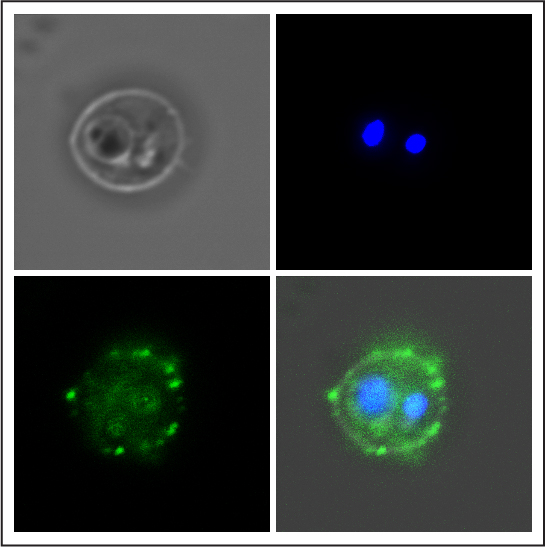
Scientists have designed a “nano-robot” made up of three DNA origami structures. © Gaëtan Bellot/Inserm
Constructing a tiny robot from DNA and using it to study cell processes invisible to the naked eye… You would be forgiven for thinking it is science fiction, but it is in fact the subject of serious research by scientists from Inserm, CNRS and Université de Montpellier at the Structural Biology Center in Montpellier[1]. This highly innovative “nano-robot” should enable closer study of the mechanical forces applied at microscopic levels, which are crucial for many biological and pathological processes. It is described in a new study published in Nature Communications.
Our cells are subject to mechanical forces exerted on a microscopic scale, triggering biological signals essential to many cell processes involved in the normal functioning of our body or in the development of diseases.
For example, the feeling of touch is partly conditional on the application of mechanical forces on specific cell receptors (the discovery of which was this year rewarded by the Nobel Prize in Physiology or Medicine).
In addition to touch, these receptors that are sensitive to mechanical forces (known as mechanoreceptors) enable the regulation of other key biological processes such as blood vessel constriction, pain perception, breathing or even the detection of sound waves in the ear, etc.
The dysfunction of this cellular mechanosensitivity is involved in many diseases – for example, cancer: cancer cells migrate within the body by sounding and constantly adapting to the mechanical properties of their microenvironment. Such adaptation is only possible because specific forces are detected by mechanoreceptors that transmit the information to the cell cytoskeleton.
At present, our knowledge of these molecular mechanisms involved in cell mechanosensitivity is still very limited. Several technologies are already available to apply controlled forces and study these mechanisms, but they have a number of limitations. In particular, they are very costly and do not allow us to study several cell receptors at a time, which makes their use very time-consuming if we want to collect a lot of data.
DNA origami structures
In order to propose an alternative, the research team led by Inserm researcher Gaëtan Bellot at the Structural Biology Center (Inserm/CNRS/Université de Montpellier) decided to use the DNA origami method. This enables the self-assembly of 3D nanostructures in a pre-defined form using the DNA molecule as construction material. Over the last ten years, the technique has allowed major advances in the field of nanotechnology.
This enabled the researchers to design a “nano-robot” composed of three DNA origami structures. Of nanometric size, it is therefore compatible with the size of a human cell. It makes it possible for the first time to apply and control a force with a resolution of 1 piconewton, namely one trillionth of a Newton – with 1 Newton corresponding to the force of a finger clicking on a pen. This is the first time that a human-made, self-assembled DNA-based object can apply force with this accuracy.

The team began by coupling the robot with a molecule that recognizes a mechanoreceptor. This made it possible to direct the robot to some of our cells and specifically apply forces to targeted mechanoreceptors localized on the surface of the cells in order to activate them.
Such a tool is very valuable for basic research, as it could be used to better understand the molecular mechanisms involved in cell mechanosensitivity and discover new cell receptors sensitive to mechanical forces. Thanks to the robot, the scientists will also be able to study more precisely at what moment, when applying force, key signaling pathways for many biological and pathological processes are activated at cell level.
“The design of a robot enabling the in vitro and in vivo application of piconewton forces meets a growing demand in the scientific community and represents a major technological advance. However, the biocompatibility of the robot can be considered both an advantage for in vivo applications but may also represent a weakness with sensitivity to enzymes that can degrade DNA. So our next step will be to study how we can modify the surface of the robot so that it is less sensitive to the action of enzymes. We will also try to find other modes of activation of our robot using, for example, a magnetic field,” emphasizes Bellot.
[1] Also contributed to this research: the Institute of Functional Genomics (CNRS/Inserm/Université de Montpellier), the Max Mousseron Biomolecules Institute (CNRS/Université de Montpellier/ENSCM), the Paul Pascal Research Center (CNRS/Université de Bordeaux) and the Physiology and Experimental Medicine: Heart-Muscles laboratory (CNRS/Inserm/Université de Montpellier).