How do bacteria manage to infect our body? What tactics do they use to slip through the cracks in our immune system? This is what the team of Thomas Henry, Inserm researcher, and his coworkers from the CNRS of Claude Bernard Lyon 1 University and the École Normale Supérieure de Lyon grouped within the International Center for Infectiology Research (CIRI) are trying to elucidate. In a paper published in Nature Communication, the researchers studied a key component in the escape mechanism of bacteria and found, in humans, the major player involved in its detection.
Detecting the presence of the enemy is the first essential step in implementing a combat response. For our bodies, that role is played by the immune system, which is confronted with various types of pathogens – particularly bacteria that employ every possible strategy to thwart its surveillance.
Normally, when invading the body, the bacteria betray their presence. The culprit, which is known as LPS and located in the bacterial cell wall, enables the human cells to recognize them and trigger an immune response. However, certain bacteria that are equipped with a more discreet form of LPS have a greater ability to pass under the immune system’s radar and increase their chances of infecting the body.
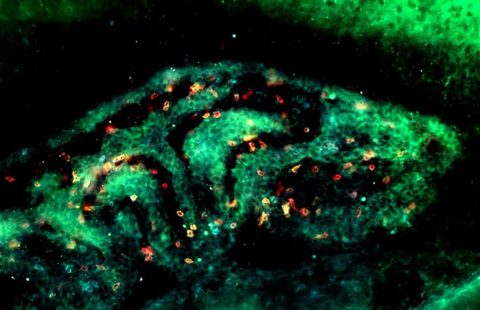
To understand the body’s defense mechanisms against bacteria, the researchers studied Francisella novicida, a pathogen model equipped with this discreet LPS. This bacterium has the capacity to escape from within the innate immune cells (macrophages) which are supposed to destroy them.
Organizing the bacterial escape
Normally, the arrival of LPS within the cytoplasm of the macrophage is detected and the inflammatory response triggered, with the death of the cell halting the propagation of the pathogen. In reality, there is a constant race between bacterial multiplication and the detection systems of the host cell. Among the macrophage’s many alarm systems, Aim2 has been identified as being that which – in the mouse – is able to detect the arrival of these bacteria within the cytoplasm. However, it is impossible to reproduce the same result in humans. So, the challenge was then to understand how this counterattack takes place in humans.
Organizing the counterattack: at the right time, together!
This discovery also helps to explain why humans are more susceptible than mice to septic shock, which occurs when bacteria invade the blood or certain organs. Given that caspase-4 is particularly sensitive, the large quantities of LPS circulating in the blood provoke an immune system surge with irreversible life-threatening consequences. Despite everything, the diversity of the detection mechanisms and their partial redundancy help ensure that, following encounters with bacteria, humans often emerge the winner.
Identification of the functioning of caspase 4 and its cofactors represents a step towards the implementation of anti-inflammatory treatments in septic shock.
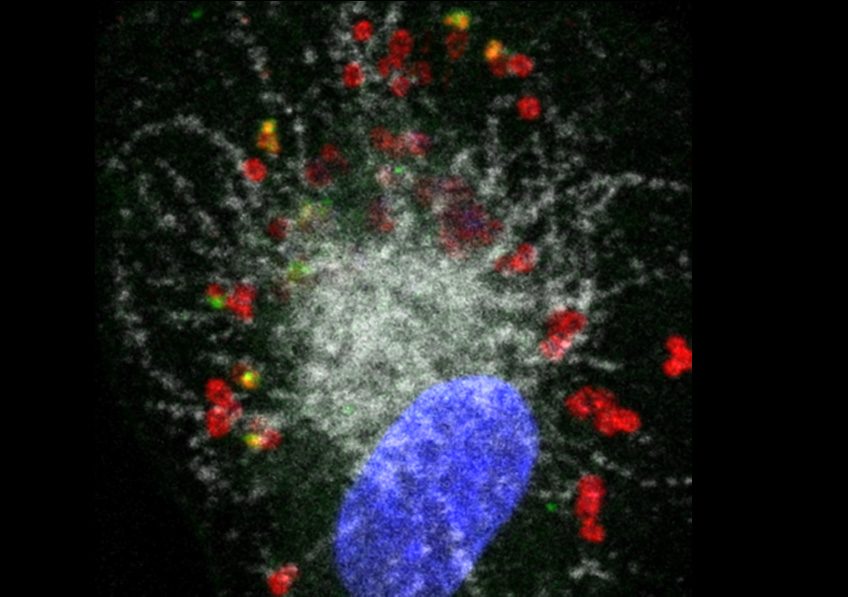
Illustration: Human macrophage (nucleus shown in blue) infected with Francisella novicida (red). The pathogen has escaped from the phagolysosomal compartment (shown in white) (one of the first defense mechanisms of the macrophage) but another defense protein, GBP2 (green), detects certain bacteria and enables caspase-4 to reveal the LPS of Francisella and implement antibacterial responses. Credit: Thomas Henry/Inserm