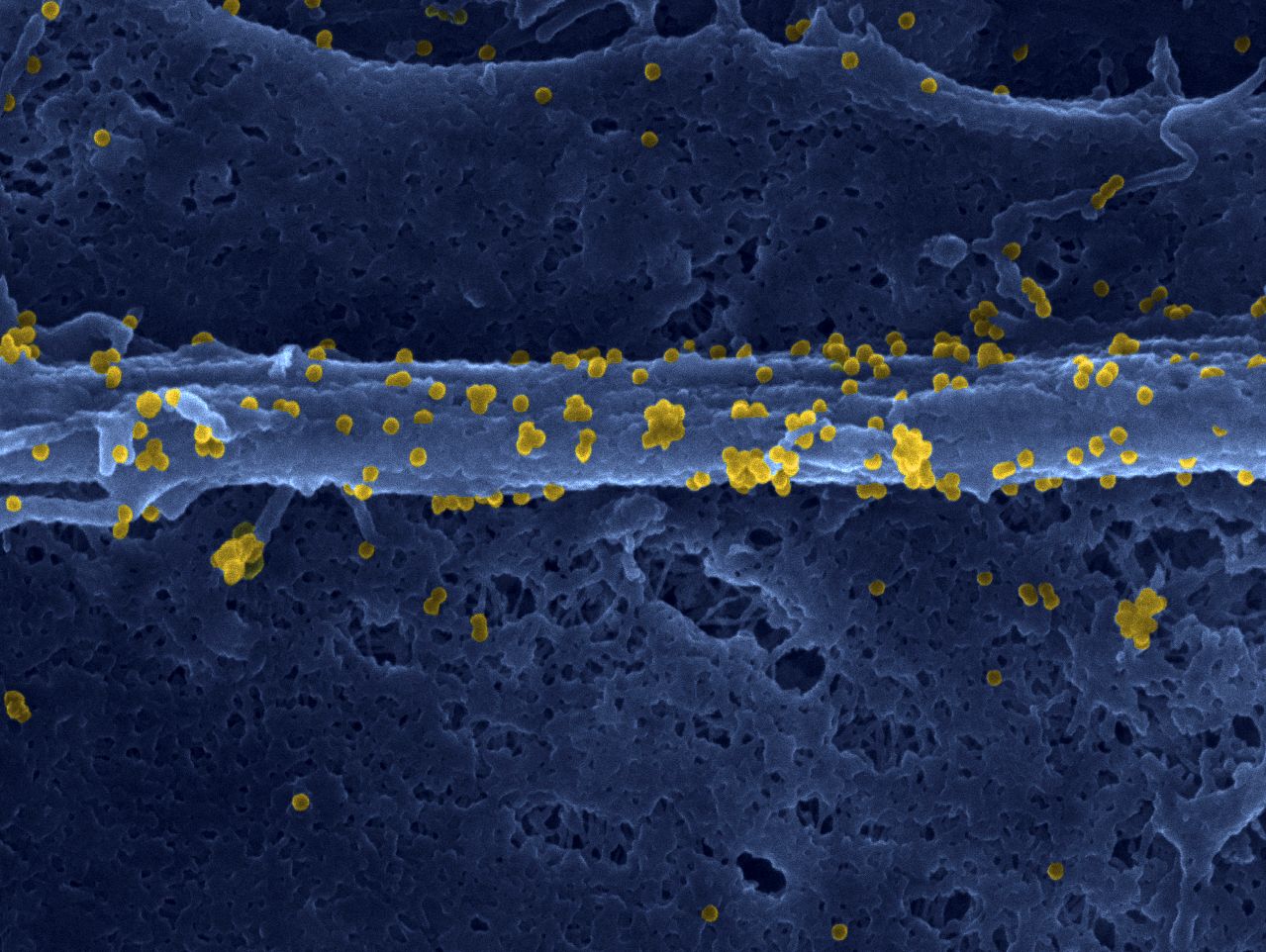
FHL1 is a major host factor for chikungunya virus infection
Laurent Meertens1,13*, Mohamed Lamine Hafirassou1, Therese Couderc2, Lucie Bonnet-Madin1, Vasiliya Kril1, Beate M. Kümmerer3, Athena Labeau1, Alexis Brugier1, Etienne Simon-Loriere4, Julien Burlaud-Gaillard5, Cécile Doyen6, Laura Pezzi7, Thibaud Goupil2, Sophia Rafasse2, Pierre-Olivier Vidalain8, Anne Bertrand Legout9, Lucie Gueneau9, Raul Juntas-Morales10, Rabah Ben Yaou9, Gisèle Bonne9, Xavier de Lamballerie7, Monsef Benkirane6, Philippe Roingeard5, Constance Delaugerre1,11, Marc Lecuit2,12 & Ali Amara1,13*
1 Inserm U944, CNRS UMR 7212, Genomes & Cell Biology of disease Unit Institut de Recherche Saint-Louis, Université de Paris, Hôpital Saint-Louis Paris France
2 Institut Pasteur, Inserm U1117, Biology of Infection Unit Paris France
3 Institute of Virology University of Bonn Medical Centre Bonn Germany
4 Institut Pasteur, G5 Evolutionary Genomics of RNA Viruses Paris France
5 Inserm U1259 MAVIVH et Plateforme IBiSA de Microscopie Electronique Université de Tours et CHRU de Tours Tours France
6 Institut de Génétique Humaine, Laboratoire de Virologie Moléculaire CNRS-Université de Montpellier Montpellier France
7 Unité des Virus Émergents, Aix-Marseille Univ, IRD190, Inserm 1207, EFS-IRBA Marseille France
8 Equipe Chimie & Biologie, Modélisation et Immunologie pour la Thérapie Université Paris Descartes, CNRS UMR 8601 Paris France
9 Center of Research in Myology, Sorbonne Université, Inserm UMRS974 Paris France
10 Département de Neurologie Centre Hospitalier Universitaire de Montpellier Montpellier France
11 Laboratoire de Virologie et Département des Maladies Infectieuses Hôpital Saint-Louis, APHP Paris France
12 Université de Paris, Department of Infectious Diseases and Tropical Medicine Necker-Enfants Malades University Hospital, APHP, Institut Imagine Paris France
13* Corresponding authors
Nature, Septembre 2019
DOI : https://doi.org/10.1038/s41586-019-1578-4