The position of cellular nuclei in muscle fibres has an important role in some muscle weaknesses. Edgar Gomes, an Inserm researcher in the myology group at the Institute of Myology (mixed Inserm/UPMC unit) recently made this discovery in collaboration with an American team. The researchers identified several proteins involved in “correctly” positioning nuclei, which is required for the muscles to function. Their results are published in a letter in the Nature review, dated 18 March.
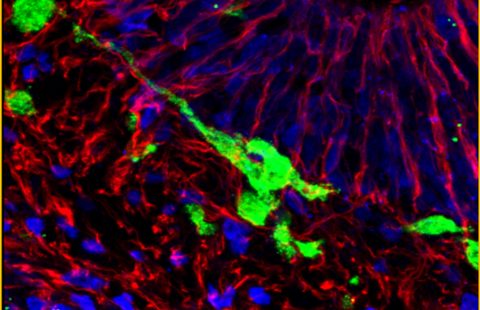
In order to move, living beings need muscles, and, more specifically, skeletal muscles that are controlled by the nervous system. Skeletal muscles are composed of cylindrical muscle fibres with a multitude of peripheral nuclei. Until now, little was known about the mechanism used to position nuclei on the edge of muscle fibres. A team of French-American researchers has tried to better understand the reasons behind nuclei layout.
Edgar Gomes and his team of collaborators have identified the mechanism involved in positioning nuclei in muscle fibres. The researchers identified (in Drosophila and mice) two proteins involved in positioning the nuclei: protein Kif5B, which belongs to the kinesin family (molecular motor), and protein MAP7, which is used to move different organelles (1) in cells.
This result was achieved by mutating MAP7 and Kif5b protein-coding genes in the Drosophila and by studying the development of the embryo. In this case, they observed that the nuclei were not correctly aligned in the muscle fibres.
“MAP7 is required to position nuclei in muscle fibre in Drosophila and in mammals” states Edgar Gomes, Inserm researcher. The research team succeeded in describing the nuclei-positioning mechanism in fibres, which involved the MAP7 protein and its interaction with the molecular motor: kinsin Kif5b. They demonstrated that a mutation of these proteins did not affect muscle extension or its attachment to the skeleton: only the position of the nuclei was affected.
By making both proteins interact together, Edgar Gomes’ team suggest that MAP7 binds with Kif5b to encourage nuclei positioning. “Furthermore, these proteins act together, both physically and genetically, and their physical bond is required for correct nuclei positioning. Our results show that they are required for the muscle to function correctly” underlines Edgar Gomes.
Muscular diseases lead to weaknesses in the fibres and can be associated with a cellular nuclei alignment failure. Edgar Gomes and his team have demonstrated that by correctly replacing the nuclei, the muscle recovers its functions. “We suggest that by correcting muscular positioning faults in patients suffering from myopathies, these patients may see improvements in their muscular functioning” concludes Edgar Gomes.
Footnote
(1) Specialized structures in the cell contained in the cytoplasm