Researcher Contact
Jean-Luc Imler03 88 41 70 36
rf.artsinu@relmi.lj
Catherine Schuster
03 68 85 37 41
rf.artsinu@retsuhcs.enirehtac
In order to infect a host cell and proliferate, some viruses, such as the hepatitis C virus, infiltrate the ribosomes, the molecular machines that assemble the proteins present in each of our cells. Viral proteins are thus produced to the detriment of cellular proteins. A group of scientists in Strasbourg has demonstrated that one of the 80 components of each ribosome is essential for infection by certain viruses without being necessary for normal cell functioning. This discovery, which may result in the development of new therapeutic strategies, was made by scientists in the Laboratoire Réponse Immunitaire et Développement chez les Insectes (CNRS) and the Institut de Recherche sur les Maladies Virales et Hépatiques (INSERM/Université de Strasbourg)1, with support from the ANRS, among others. It is the subject of an article published in Cell on 20 November 2014.
A viral infection can be treated by blocking certain viral components. However, these are far less numerous than the host cellular proteins with which they interact. Furthermore, these factors mutate much more easily and can therefore escape treatment. For these reasons, virologists are seeking to develop antiviral agents that can target these cellular proteins (or factors). But there is one downside, and it is considerable: the factors targeted by this strategy often play a crucial role in the cell, causing adverse effects.
In this context, a group of scientists in Strasbourg (France) has identified a promising cell component called RACK1 in the ribosome, the complex cellular machinery where proteins are assembled. RACK1 could become the target for new types of antiviral therapies as it has been shown to be necessary for the infection of cells by certain viruses — but not essential for normal cell functioning.
The ribosome is an assembly line for proteins, made of amino acids whose alignment is determined by the genetic message the ribosome can read (contained in messenger RNA molecules). The strategy deployed by numerous viruses in order to replicate consists in entering the ribosome of an infected cell, thus forcing the manufacture of their own proteins, to the detriment of cellular proteins. This leads to the production of new viral particles that can infect other cells, and so on. The present study has shown that among the 80 or so sub-units that make up the ribosome, RACK1 is a portal of entry for several viruses, including hepatitis C. More remarkable still is the fact that most cellular mRNA can be translated into proteins in a RACK1-depleted ribosome, while this sub-unit is essential for the RNA translation — and hence the replication— of certain viruses.
The scientists made this discovery by working on the fruit fly (Drosophila melanogaster). RACK1 -depleted adult flies survived normally but could no longer be infected by certain insect viruses. The same observation was made relative to human cells in culture:the absence of RACK1 did not compromise their survival or replication, but prevented infection by the hepatitis C virus. And this may be valid for other viruses that adopt the same cell piracy strategy (such as polio and foot-and-mouth viruses or enteroviruses, etc.).
This discovery thus opens the way towards new therapeutic opportunities based on inhibiting this viral junction point on the cell ribosome. The fact that this mechanism can be used by viruses of markedly different types suggests that it may be possible to develop treatments with a broad spectrum of action that can be used to combat viral infections in insects, animals and humans.
However, if the RACK1 protein is conserved in species as different as the fruit fly and humans, it is probably because it has a function in these organisms. Indeed, although the adults are viable, RACK1- depleted fruit fly larvae and mouse embryos do not survive beyond a certain developmental stage. This means that cellular mRNA deployed in specific situations require RACK1 for their translation. Identifying the conditions under which RACK1 is useful to cells will therefore be crucial before it can be used as a therapeutic target.
At a fundamental level, these findings show that the translation of RNAs into proteins is more complex than previously thought. But they provide opportunities to elucidate the “ribosomal code” (superimposed on the genetic code and on other mechanisms that regulate gene expression); depending on the composition and structure of the ribosome, some RNAs may be selectively translated, and others not. The clues in favor of such a code are accumulating, but it now needs to be deciphered.
This work received support notably from the ANRS (France REcherche Nord&sud Sida – Hiv Hépatites), the FRM (Fondation pour la Recherche Médicale), the Fondation ARC for cancer research and the Institut Hospitalo-Universitaire de Strasbourg Mix-Surg.
1) in collaboration with the Laboratoire Architecture et Réactivité de l’ARN (CNRS) and the Laboratoire Spectrométrie de Masse Biologique et Protéomique (CNRS/ESPCI ParisTech).
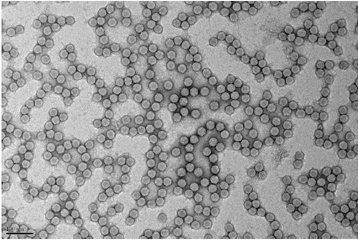
>Preparation of the DCV (Drosophila C virus) used during the study. © Jean-Luc Imler<