Credits: Fotolia
In a paper published in the journal Science Translational Medicine , Guy and his team Gorochov research center CIMI (Inserm / Université Sorbonne) and Immunology Department at the Pitié-Salpêtrière Hospital, AP-HP, reveal that our IgA act as conductor of the intestinal microbiota.They effectively prevent intestinal colonization by the oral flora and promote the presence of certain bacteria, totally innocent of infectious standpoint, but playing a beneficial role.
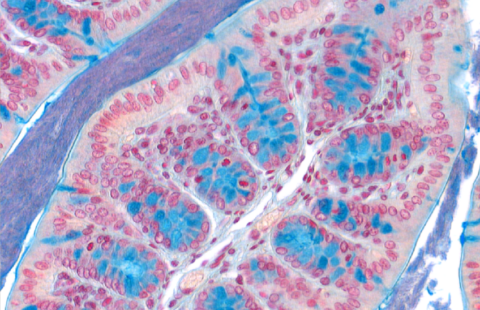
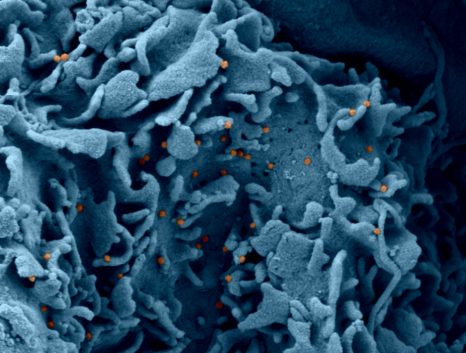
Our pact with microbes, otherwise known as symbiosis, we make them indispensable to normal life. Obesity, cancer, autoimmunity, accompanied unlike dysbiosis, that is to say, a disturbance of the bacterial ecosystem in favor of the action of “bad” bacteria. Until recently, the IgA antibodies that we secrete heavily in our digestive tract (66 mg / kg / day) was considered a defense to prevent the passage of potentially harmful bacteria through the intestinal barrier so that its effects potential microbial ecology sheltered by man remained unclear. This is precisely what the researchers wanted to understand.
It is not possible to inactivate a gene in humans to elucidate its function, as is done in mice. To assess the impact of IgA on the microbiota, the authors have taken advantage of a clinical situation of immune deficiency resulting in the almost complete absence of IgA in blood and secretions. Typical bacterial targets of IgA in the general population were also determined by purifying the part of the fecal microbiota naturally covered with IgA in healthy subjects, a unique approach developed by Martin Larsen in the laboratory. Then, total or fractionated microbiota were analyzed in a so-called metagenomic approach comprising sequence simultaneously all bacterial genomes present in a sample. Finally,
The work published today reveals that IgA plays an organizing role of the intestinal microbiota. IgA prevent intestinal colonization by the oral flora while promoting the presence of certain commensal, totally innocent of an infectious standpoint, but play a beneficial role.
This work has also helped to break an old mystery by explaining why the IgA deficiency (affecting about 1 in 500 Caucasian) is not accompanied more often fatal infections. The study shows that IgM, another type of antibody, can partly compensate IgA in its functions of interaction with the microbiota. A compensation however incomplete because patients with IgA deficiency suffer from respiratory infections, but also autoimmunity and atopy. These symptoms although point out the specific roles and not strictly anti-infectives, played by IgA.
These findings were obtained with the help of 21 deficient patients IgA, followed in hospitals of the AP-HP. Besides the fundamental breakthrough in understanding the role of IgA in the establishment of a physiological balance essential to health, the article opens up new therapeutic prospects oral supplementation these IgA deficient patients.
Finally, this study illustrates how anti-microbiota analysis of the antibody response can be a convenient way to study the interface between the host and its microbiota own, and thus the immune footprint of that scale the entire body. The study of anti-microbiota individual serological signatures representing a new biomarker for studying microbiota associations / disease that currently show the open, especially in the field of cancer.