Researcher Contact
Yamina Bennasser
Chargée de recherche Inserm
Equipe « Virologie Moléculaire »
UMR9002, Institut de génétique humaine (IGH)
+33 (0)4 34 35 99 88
Press Contact
rf.mresni@esserp
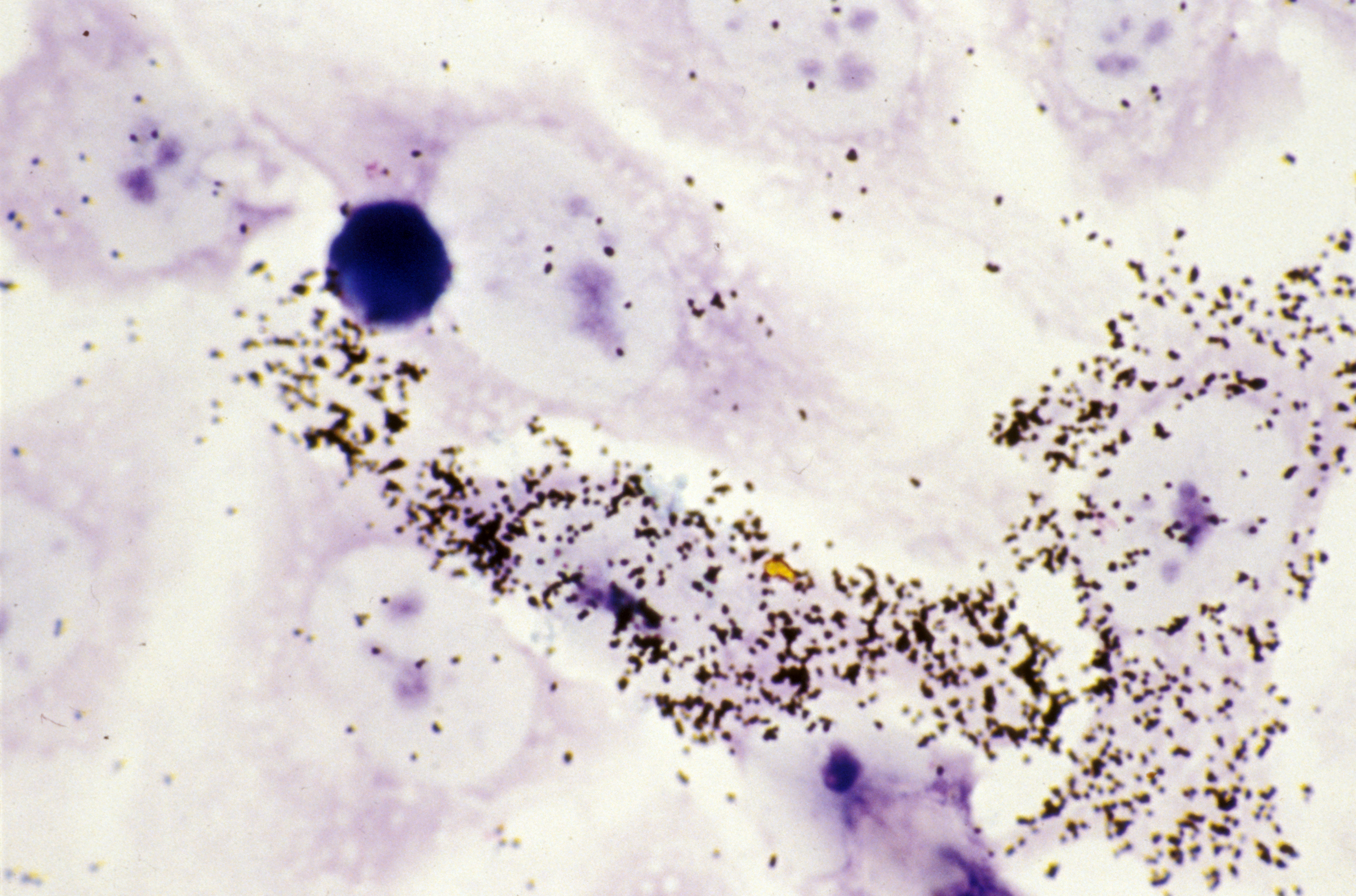
Hybridation in situ des ARN du virus du sida (VIH) dans les cellules immunitaires infectées (HeLa CD4+). ©Inserm/Fournier, Jean-Guy, 1994
How is HIV able to escape the surveillance of the immune system within the very cells it infects? Researchers from Inserm, CNRS, Université de Montpellier and Université de Lorraine decided to take a closer look at one of these evasion mechanisms. They were able to observe the ability of HIV to “camouflage” its RNA within the infected cell using an intracellular enzyme. This research, published in Nature, provides new knowledge on the evasion mechanisms of HIV in the face of the innate immune system.
From the very first stages of a viral infection, the intracellular “radars” of the innate immune system make it possible to rapidly trigger an antiviral response through the secretion of type I interferons, proteins produced by the white blood cells to regulate and stimulate the immune response.
The Human Immunodeficiency Virus (HIV) targets the cells of the immune system and causes severe immune deficiencies responsible for AIDS. When HIV infects a cell, its genome made up of single-stranded RNA is transformed into DNA. It then inserts itself into the nucleus of the host cell where it incorporates into its genome. The success of these early stages depends on the ability of the virus to camouflage itself within the cell and pass unnoticed by its detectors, particularly those capable of recognizing the nucleic acids of its genome as foreign RNA.
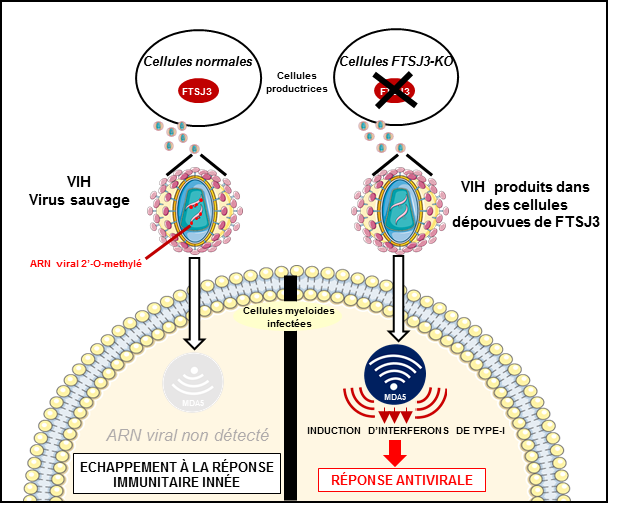
Researchers from Inserm, CNRS, Université de Montpellier and Université de Lorraine decided to take a closer look at this mechanism through which HIV can evade the surveillance of the cells by using a system of camouflage. Within the cells, an enzyme called FTSJ3 is found which is capable of modifying some of the nucleic acids comprising cell RNA by adding to them a methyl group. This modification is a signature of self (set of molecules resulting from the expression of the individual’s genome, as opposed to non-self) which enables the detectors to recognize the cellular RNA as such in the human cells and to avoid its destruction by the immune system.
These findings represent a significant advance in the understanding of HIV infection by revealing a new evasion strategy of the virus when confronted with the innate immune system’s cellular detection system. Further elucidation of these evasion mechanisms could in the longer term enable the development of therapeutic and/or vaccine strategies aimed at modifying the virus so that it leads to the establishment of an antiviral response which, when this is early, enables the cell to implement an immune response and control the infection.
Yamina Bennasser
Chargée de recherche Inserm
Equipe « Virologie Moléculaire »
UMR9002, Institut de génétique humaine (IGH)
+33 (0)4 34 35 99 88
rf.mresni@esserp
FTSJ3 is an RNA 2′-O-methyltransferase recruited by HIV to avoid innate immune sensing
Mathieu Ringeard1, Virginie Marchand2, Etienne Decroly3, Yuri Motorin4 & Yamina Bennasser1
1 IGH, CNRS, Université de Montpellier, Montpellier, France
2 Next-Generation Sequencing Core Facility, UMS2008 IBSLor CNRS-University of Lorraine-INSERM, BioPole, Vandoeuvre-les-Nancy, France
3 CNRS, Aix-Marseille University, AFMB, Marseille, France
4 IMoPA UMR7365 CNRS-University of Lorraine, BioPole, Vandoeuvre-les-Nancy, France