Researcher Contact
Stanislas DehaeneProfesseur au Collège de France
Directeur de l’Unité Inserm 992 « Neuroimagerie cognitive » (Inserm/CEA/Université Paris-Sud)
rf.aec@eneahed.salsinats
Researchers from the Cognitive Neuroimaging Unit at NeuroSpin have just identified a network of areas of the brain that are organised in a way that could at least partially explain the specificity of the cognitive functions in the human species. These regions are specifically activated in humans, but not in the macaque monkey, in response to specific variations in the auditory sequences played. They coincide with the classic language areas, particularly Broca’s area. The human faculty of language could therefore have its origins in the emergence of a brain circuit capable of combining, in a single region, information coming from the other regions of the brain into a coherent whole. These results, obtained through a collaboration between the French Atomic Energy Commission (CEA), Inserm, Collège de France, Versailles-Saint-Quentin-en-Yvelines University and Paris-Sud University, are published in Current Biology.
In this study, conducted at NeuroSpin, Stanislas Dehaene (a professor at Collège de France, and director of the Inserm/CEA/Paris-Sud University Cognitive Neuroimaging Unit) and Bechir Jarraya (Professor of Neurosurgery at Versailles-Saint-Quentin-en-Yvelines University), together with Liping Wang and Lynn Uhrigh, used a noninvasive functional imaging method, 3 Tesla functional MRI. They exposed three macaque monkeys and twenty volunteers to regular auditory sequences, e.g. three identical sounds followed by a fourth different sound (a sequence notated as AAAB). Occasionally, they presented a sequence that violated this regularity, either because it included a different number of sounds (e.g. AAAAAB), or because the sequence of sounds was abnormal (e.g. AAAA, which does not end in a B sound).
The monkey’s brain reacted to changes in numbers and sequences, which denotes a certain capacity for abstraction. However, it did so in distinct areas, specialised for either number or sequence. In contrast, the human brain combined the two parameters in regions that coincide with the language areas.
Thus, whereas the monkeys detected isolated properties, such as “four sounds” or “the last one is different,” evolution seems to have endowed our species with a specific ability to combine these pieces of information into a coherent whole, a formula such as “three sounds, then another”—the very beginnings of an inner language?
Thus, although the abstract representation of sound sequences is possible in non-human primates, the evolution of a new brain circuit, connected to the auditory areas, might have enabled our species to acquire the unique ability to compose and recognise the complex sequences characteristic of human languages.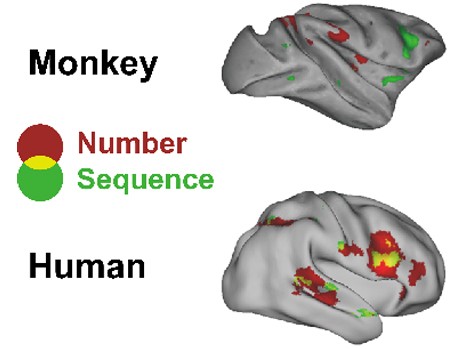
This figure illustrates the unique ability of the human brain to combine pieces of abstract auditory information. Some regions of the brain are associated with the detection of a change in number of sounds by the brain, independently of a concomitant change in the sequence of sounds (red areas in the figure). Conversely, some regions of the brain detect changes in the sequence of sounds, independently of their number (green areas). In the monkey brain, these two sets of regions are unconnected. Places where they intersect (shown in yellow), i.e. regions that combine the two types of information, “change in sequence of sounds” and “change in number of sounds,” are found only in the human brain. All activations detected are projected on a lateral view of the right hemisphere for purposes of representation. © Liping Wang
Representation of Numerical and Sequential Patterns in Macaque and Human Brains
Wang L¹, Uhrig L², Jarraya B³, Dehaene S4. 1 Cognitive Neuroimaging Unit, INSERM, 91191 Gif-sur-Yvette, France; CEA, DSV, I2BM, NeuroSpin Center, 91191 Gif-sur-Yvette, France; Key Laboratory of Brain Functional Genomics (MOE and STCSM), Institute of Cognitive Neuroscience, School of Psychology and Cognitive Science, East China Normal University, 200062 Shanghai, China; NYU-ECNU Institute of Brain and Cognitive Science, NYU Shanghai, 200062 Shanghai, China. Electronic address: moc.liamg@gnwgnipil. 2 Cognitive Neuroimaging Unit, INSERM, 91191 Gif-sur-Yvette, France; CEA, DSV, I2BM, NeuroSpin Center, 91191 Gif-sur-Yvette, France; Inserm Avenir-Bettencourt-Schueller Team, 91191 Gif-sur-Yvette, France. 3 Cognitive Neuroimaging Unit, INSERM, 91191 Gif-sur-Yvette, France; CEA, DSV, I2BM, NeuroSpin Center, 91191 Gif-sur-Yvette, France; Inserm Avenir-Bettencourt-Schueller Team, 91191 Gif-sur-Yvette, France; Université Versailles Saint-Quentin-en-Yvelines, 78000 Versailles, France; Neuromodulation Unit, Department of Neurosurgery, Foch Hospital, 92150 Suresnes, France. 4 Cognitive Neuroimaging Unit, INSERM, 91191 Gif-sur-Yvette, France; CEA, DSV, I2BM, NeuroSpin Center, 91191 Gif-sur-Yvette, France; Collège de France, 75005 Paris, France; Université Paris-Sud, 91400 Orsay, France.
Current Biology, 30 juillet 2015