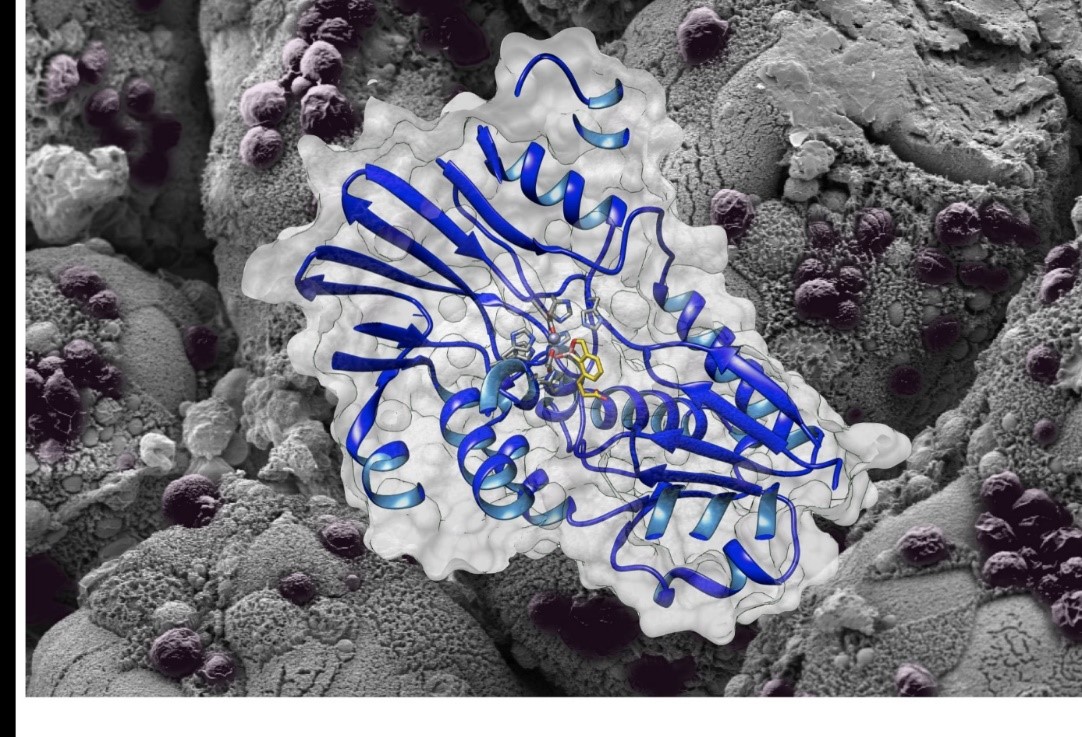
3D structure of the enzyme with the molecule AN3661 shown against the background of the intestine of an immunocompromised mouse infected with Cryptosporidium. ©Fabrice Laurent and Christopher Swale.
Researchers from Inserm and INRA working in the teams of Mohamed-Ali Hakimi (Institute for Advanced Biosciences – Inserm U 1209 / CNRS JRU 5309 / UGA) and Fabrice Laurent (INRA) have recently discovered a new candidate drug to control cryptosporidiosis, a severe intestinal disease in children, immunocompromised patients, and young ruminants. Beyond this disease, their research represents an opportunity to discover new therapeutic avenues for related infections, such as toxoplasmosis and malaria. Their findings have been published in Science Translational Medicine.
Cryptosporidiosis is a diarrheal disease caused by Cryptosporidium, a microscopic parasite that develops in the intestine of numerous mammals – notably humans. This intestinal parasite is mainly spread through contaminated drinking or pool water, where it can survive for several days in the presence of chlorine, or through contact with infected animals. Over the past 20 years, infection with Cryptosporidium has been recognized as a common cause of waterborne disease in humans. According to a recent study by the US Centers for Disease Control and Prevention (CDC), the number of Cryptosporidium epidemics is even on the increase. In humans, it causes acute and sometimes fatal diarrhea in the most vulnerable populations, including young malnourished children and immunocompromised patients (for example, those infected with HIV). The therapeutic arsenal is currently very limited and in some cases ineffective in eliminating this parasite.
This study conducted by the researchers from Inserm and INRA reveals the discovery of a candidate drug called AN3661, which drastically reduces not just Cryptosporidium infection but also that of Toxoplasma, the parasite responsible for toxoplasmosis.
The study teams have revealed the mechanism of action of this molecule by elucidating the three-dimensional structure of its target, called CPSF3, in Cryptosporidium. AN3661 binds to the heart of the enzyme CPSF3, thereby preventing the maturation of the messenger RNA, a process essential to the parasite’s survival. Preclinical tests using an animal model show remarkable efficacy in vivo with single-dose treatments of the infection in immunocompromised or baby mice.This major discovery paves the way for new therapeutic strategies and innovations to fight not just cryptosporidiosis but also other related infections, such as toxoplasmosis and malaria.
These contents could be interesting :
Metal-Captured Inhibition of pre-mRNA Processing Activity by CPSF3 controls very efficiently Cryptosporidium Infection. Christopher Swale1, Alexandre Bougdour1, Audrey Gnahoui-David2, Julie Tottey2, Sonia Georgeault3, Fabrice Laurent2*, Andrés Palencia1,4*†, Mohamed-Ali Hakimi1*† 1Institute for Advanced Biosciences (IAB), Host-Pathogen Interactions and Immunity to Infection, INSERM U1209, CNRS UMR 5309, University Grenoble Alpes, 38000 Grenoble, France. 2 INRA, Université François Rabelais de Tours, Centre Val de Loire, UMR1282 ISP, Laboratoire Apicomplexes et Immunité Mucosale, 37380 Nouzilly, France. 3Plateforme des Microscopies, Université et CHRU de Tours, 37000 Tours, France. 4Institute for Advanced Biosciences (IAB), Structural Biology of Novel Drug Targets in Human Diseases, INSERM U1209, CNRS UMR 5309, University Grenoble Alpes, 38000 Grenoble, France. *Corresponding author. Email: rf.mresni@imikah.ila-demahom (M.-A.H.); andres.palencia@univ- grenoble-alpes.fr (A.P.); rf.arni@tnerual.ecirbaf (F.L.) †These authors contributed equally to this work. Science Translational Medicine https://dx.doi.org/10.1126/scitranslmed.aax7161