An international team, including researchers in France at Inserm, CNRS and the University of Strasbourg, brought together at IGBMC[1] is lifting the veil on the molecular mechanisms causing heart dysfunctions in myotonic dystrophy, a genetic disease affecting one person in 8,000. This new study, published this week in Nature Communications, could contribute to discovering a treatment.
(c) Inserm/IGBMC
Myotonic dystrophy, also known as Steinert disease, is the commonest adult form of muscular dystrophy. Patients affected by this genetic condition suffer from wasting of skeletal muscles as well as arrhythmia and other cardiac dysfunctions. This is a particularly debilitating disease, for which there is currently no treatment.
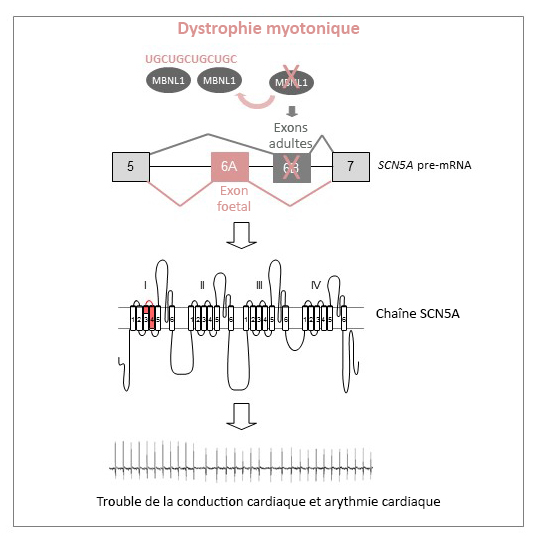
Myotonic dystrophy is due to a mutation leading to the expression of RNA containing long repetitive sequences of the CUG trinucleotide. These mutated RNAs accumulate and alter regulation of alternative splicing[2] of numerous genes. Despite the significance of work already done on this disease, many points remain to be elucidated. This is true for the origin of arrhythmia and other cardiac dysfunctions, which represent the second most common cause of death in this disease.
In this new study, researchers have identified new splicing alterations in messenger RNA from heart samples of affected patients. Among these many alterations, biologists have established that those relating to the cardiac sodium channel (SCN5A) were fundamental to understanding the cardiac dysfunctions of these patients.
Scientists there clarified the molecular mechanisms leading to the alteration of SCN5A in these patients. Collaboration with Denis Furling’s team at the Institut de Myologie in Paris has enabled these cardiac alterations to be reproduced in a mouse model.
“The next step would be to see if, by restoring correct splicing of SCN5A, we can also successfully restore normal heart function”, explains Nicolas Charlet-Berguerand, Inserm Research Director, who coordinated this work. The researchers are hoping that this breakthrough will give a fresh boost to research into this rare disease.
Alternative splicing model of the cardiac sodium channel (SCN5A) in myotonic dystrophy. (c) Inserm/IGBMC
This work was financed by the French Myopathy Association (AFM), the European research council (ERC), the European E-rare programme (ANR), Inserm and Labex-INRT (ANR).
[1] Institute of Genetics and Molecular and Cellular Biology (Inserm/CNRS/University of Strasbourg)
[2] In eukaryotes, this is a process by which RNA transcribed from a gene can undergo different cutting and splicing steps leading to the loss of various regions. This process enables proteins having distinct properties to be produced from the same gene.