While most of us recover from influenza after a week, it can be a very severe disease, and even fatal in rare cases, with no reason for physicians to have expected such an outcome. By analysing the genome of a little girl who contracted a severe form of influenza at the age of two and a half years, researchers at the Laboratory of Human Genetics of Infectious Diseases (a joint French-American international laboratory), which brings together researchers from Inserm, Paris Descartes University, and physicians from the Paris public hospitals (AP-HP; Necker Hospital for Sick Children), working at the Imagine Institute, and from The Rockefeller University in New York, have discovered that she has a genetic mutation, unknown until now, that causes a subtle dysfunction in her immune system. More generally, these results show that genetic mutations could be the root cause of some severe forms of influenza in children, and indicate in any event that immune mechanisms missing in this little girl are needed for protection against this virus in humans. These results are published in the journal Science Express. ©fotolia
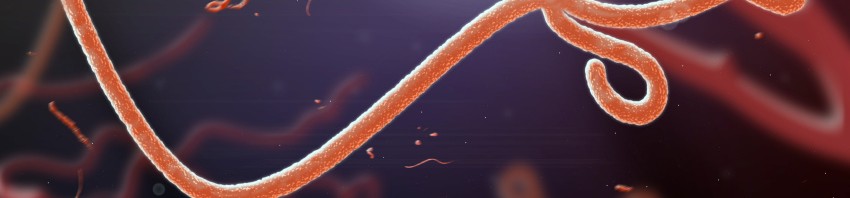
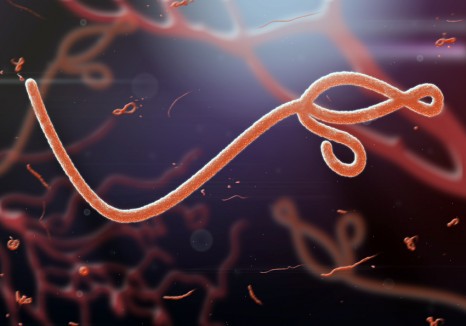
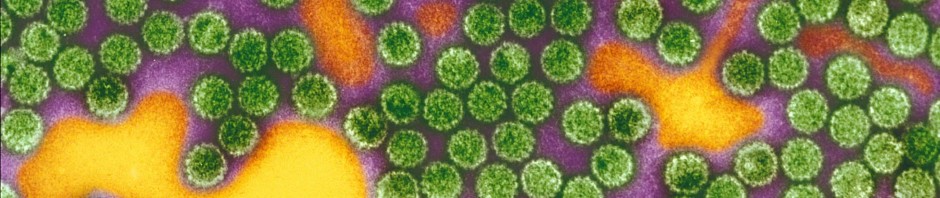
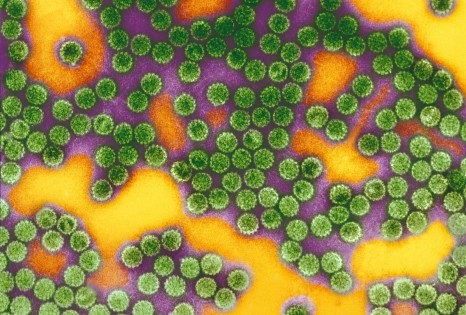
Seasonal influenza is an acute viral infection caused by the influenza virus. It is characterised by high fever, headaches, sore muscles, etc. Apart from vaccination, there is no treatment for it other than symptomatic (pain) treatment. In most cases, patients recover after a week, but in more vulnerable people influenza can cause acute respiratory distress, which is potentially fatal.
The main known risk factors for severe forms of influenza are some acquired comorbidities, such as chronic lung disease. However, the cause of most fatal cases remains unexplained, especially in children.
The absence of cases of severe influenza in patients with known acquired immunodeficiencies, which usually increase susceptibility to infections, is also surprising.
Given these different observations, the researchers at Jean-Laurent Cassanova’s and Laurent Abel’s laboratory, in Paris and New York, therefore formulated a hypothesis whereby severe influenza in healthy children might be the result of genetic errors.
To test this hypothesis, they sequenced the entire genome of a 7-year-old child who had contracted a severe form of influenza (influenza A virus strain H1N1), requiring her admission to a paediatric intensive care unit in January 2011, at the age of two and a half years. At the time, she showed no other known pathology that might have suggested greater vulnerability to the virus than that of other children.
This analysis, combined with analysis of her parents’ genomes, made it possible to show that the little girl had inherited a mutated allele of the gene encoding interferon regulatory factor (IRF7) from both of her parents. The latter is a transcription factor known to amplify the production of interferons in response to viral infection in mice and humans.
In contrast to her parents, in whom the mutation of a single allele of the gene is of no consequence, in the little girl, mutation of both alleles of the gene encoding IRF7 has led to its inactivation. The result: failure to produce interferons, disrupting her system of defence against influenza virus infection in a cascading manner.
By carrying out a comprehensive series of experiments on blood cells, particularly dendritic cells, and by generating lung cells from stem cells taken from the young girl, the researchers provided proof that the mutations observed in this little girl explain the development of severe influenza. Furthermore, this discovery demonstrates that interferon amplification dependent on IRF7 expression is needed for protection against influenza virus in humans. They now need to search for mutations in this or other genes in other children recruited following an episode of unexplained severe influenza.
Based on these initial observations, the researchers at Inserm believe that therapeutic strategies based on recombinant interferons, available in the pharmacopoeia, could help to combat severe forms of influenza in children.
This multidisciplinary study required multiple collaborations in Europe and the United States.