© L. Peris /GIN
After 40 years of research, researchers at the CEA, the CNRS, the University of Grenoble-Alps, the University of Montpellier and the Inserm have finally identified the enzyme responsible for the tubulin cycle. Surprisingly, it is not one enzyme but two which control the cycle of this essential component of the cytoskeletal structure. This work opens up new prospects for the improved understanding of the role of tubulin, changes in the cycle of which are associated with cancers, cardiac diseases and neural disorders. These results were published on 16th November 2017 in the review Science.
A collaborative international project involving researchers from the CEA (French Atomic Energy Commission), the CNRS (National Centre for Scientific Research), the Inserm (French National Institute of Health and Medical Research), the University of Grenoble-Alps, the University of Montpellier and the University of Stanford[1] has identified an enzyme, Tubulin CarboxyPeptidase (TCP), which is responsible for the biochemical transformation of cellular microtubules, or detyrosination. Detyrosination is a biological reaction for the removal of the terminal amino acid tyrosine[2] from tubulin α, a constituent of microtubules. After four decades of research, biologists have succeeded in isolating this protein by purification, and have gone on to provide evidence of its cellular activity.
Microtubules contribute to essential cellular functions
Microtubules are dynamic fibres which are present in all cells. Formed by the combination of two proteins (tubulin α et tubulin β), microtubules assume numerous functions. They separate the chromosomes which are to be contained in the two daughter cells resulting from cell division, they contribute to the polarity of cells, morphology and cellular migration. They form “rails” upon which cellular constituents, such as proteins or RNA strands, are transported.
These cellular functions are regulated by the existence of “signals” which are present on the surface of microtubules. These signals are biochemical modifications to amino acids (described as post-translational modifications, as they take place after protein synthesis), executed by various enzymes which, in this case, modify the tubulins.
The enzyme TCP, identified after 40 years of mystery
The activity of one of these enzymes was identified for the first time in 1977 by Argentine researchers, who named it “TCP” (Tubulin CarboxyPeptidase). The function of this enzyme, which had never been identified previously (its size and sequence were unknown) is the removal of the terminal amino acid, a tyrosine, from the end of tubulin α. This is the detyrosination reaction. A reverse enzyme, ligase TTL, is responsible for resetting this tyrosine in its place. This is tyrosination. This detyrosination/tyrosination cycle is vital for the cell and the organism. Massive (abnormal) detyrosination is observed in a number of severe cancers and cardiac diseases.
The identification and characteristic definition of TCP was therefore a major objective for understanding the physiological function of the detyrosination of tubulin α and evaluating the consequences of its inhibition.
In order to isolate TCP, researchers have monitored its activity, employing conventional biochemical techniques, and have involved chemists from the University of Stanford, who have developed a small inhibitor molecule for its activity. This molecule has been used as bait to “reel in” the desired enzyme.

Tubulin detyrosination/tyrosination cycle
Microtubules are fibres which are present in all cells, comprised of a stack of α/β tubulins. Tubulin carries a tyrosine (Y) at its end, which is alternately removed and replaced by two enzymes, thereby modifying the surface of microtubules. TCP (which is represented by a saw comprised of two elements, VASH/SVBP) is responsible for detyrosination. TTL (represented by a tube of glue) resets tyrosine on the tubulin. This cycle is essential to the various functions of microtubules in cells (division, migration, etc.) and is vital for the organism. © C. Bosc, GIN
| SVBP | SVBP |
| VASH1,2 | VASH1,2 |
| scie TCP | TCP saw |
| detyrosination | detyrosination |
| tubuline tyrosinée | tyrosinated tubulin |
| tubuline détyrosinée | detyrosinated tubulin |
| tyrosination | tyrosination |
| colle TTL | TTL glue |
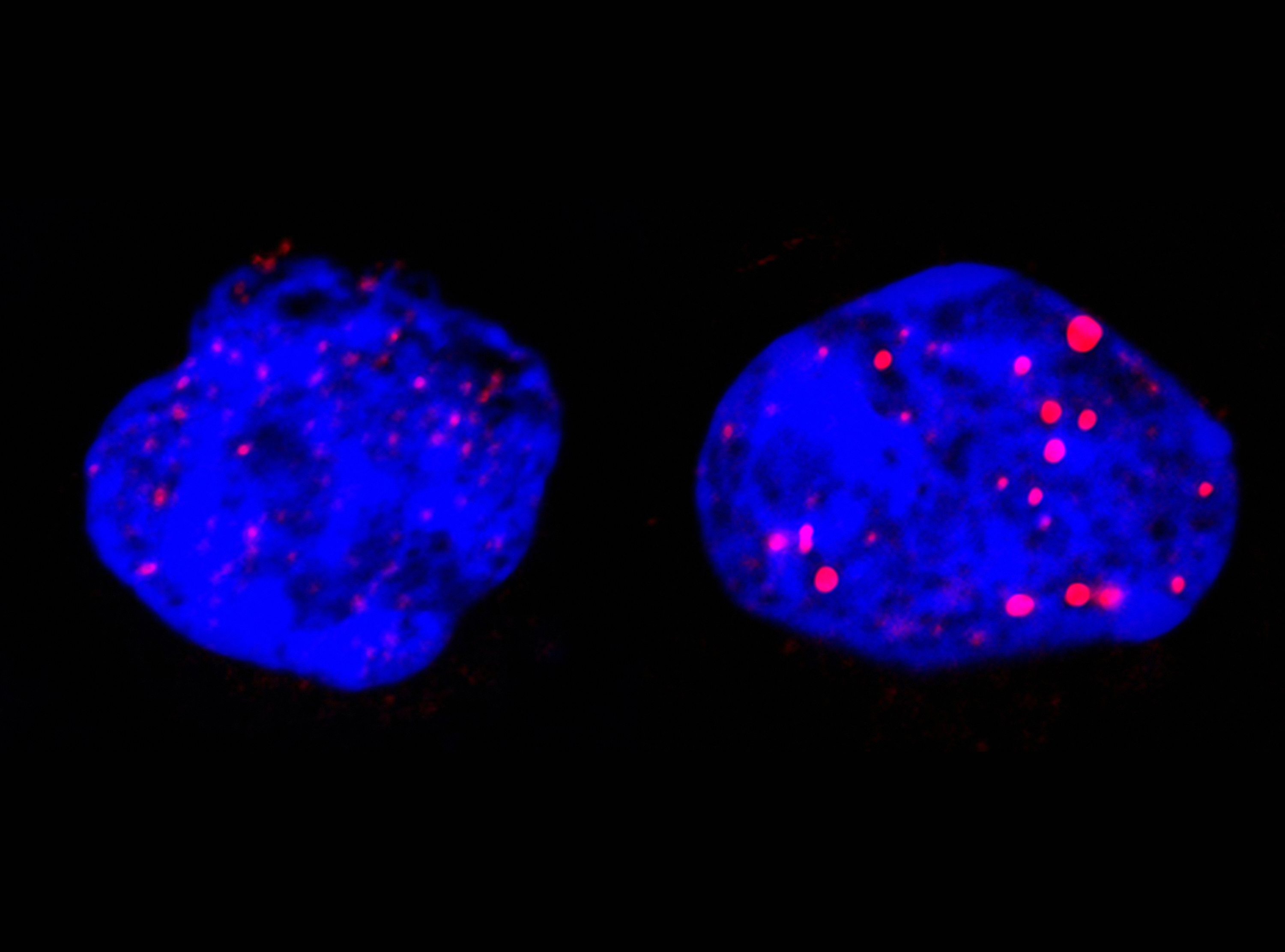
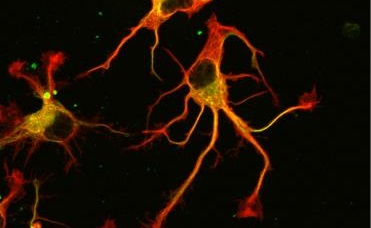
Ultimately, not one, but two enzymes have been discovered. The latter, named VASH1 and VASH2, were already known to scientists, but it was not known that these were enzymes associated with the cytoskeleton. Researchers have demonstrated that, provided they are associated with a partner protein called SVBP, VASH1 and VASH2 are capable of the detyrosination of tubulin α. To demonstrate this, researchers have inhibited the expression of the former (or that of their partner SVBP) in neurons. They then observed a very strong decline in the rate of detyrosination of tubulin α, together with anomalies in the morphology of neurons (see Figure). Researchers went further, demonstrating that these enzymes are also involved in the development of the cerebral cortex.
Prospects for the fight against cancer
Thus, forty years after the conduct of the first work on the detyrosination of tubulin α, the enzymes responsible have been revealed. Scientists are now hoping that, by modulating the effectiveness of TCP and improving their knowledge of the detyrosination/tyrosination cycle, they can advance the fight against certain cancers, and achieve progress in the understanding of cerebral and cardiac functions.

| Contrôle | Control |
| VASH1 et VASH2 réduites | VASH1 and VASH2 reduced |
| SVBP réduite | SVBP reduced |
| Tubuline deTyrosinée / Tubuline Tyrosinée | Detyrosinated tubulin / Tyrosinated tubulin |
Photographs of the alteration of neurons associated with a reduction in the expression of TCP enzymes (VASH/SVBP). From left to right: control neuron, neurons in which the expression of VASH1 and VASH2 is reduced, neurons in which the expression of SVBP is reduced. Neurons with a reduced enzyme show a delay in development, together with morphological anomalies.
[1] The following institutes are involved: Grenoble Institute of Neurosciences, GIN (Inserm/Univ. Grenoble-Alps); Institute of Biosciences and Biotechnologies of Grenoble, BIG (Inserm/CEA/Univ. Grenoble-Alps); Institute of Advanced Biosciences, IAB (Inserm/CNRS/Univ. Grenoble-Alps), Department of Pathology, Stanford University School of Medicine (Stanford, USA), Institute of Human Genetics, IGH (CNRS/Univ. of Montpellier), Montpellier Centre of Cell Biology Research, CRBM (CNRS/Univ. of Montpellier).
[2] Tyrosine is one of the 22 constituent amino acids in proteins.