Inserm teams led by Prof. Jean-Yves Blay and Christophe Caux in Lyon[1], and by Franck Tirode and Olivier Delattre in Paris[2] have just demonstrated a new genetic variant in tumours that had not been identified until now. Their results enable better diagnosis of these tumours using a validated biomarker. This study is published in the journal Nature Genetics.
The application of so-called “high-throughput” sequencing techniques to oncology is currently transforming our understanding of the genesis and natural progression of cancers. These new data have partly called into question the tumour classifications based on traditional techniques such as pathology. According to Prof. Jean-Yves Blay, codirector of this study, “Our priority is to refine tumour classifications used in clinical practice in order to offer patients the most appropriate therapeutic option. This is particularly important where sarcomas are concerned, where we are regularly faced with the problem of highly undifferentiated tumours that are difficult to classify.”
A single objective, two strategies
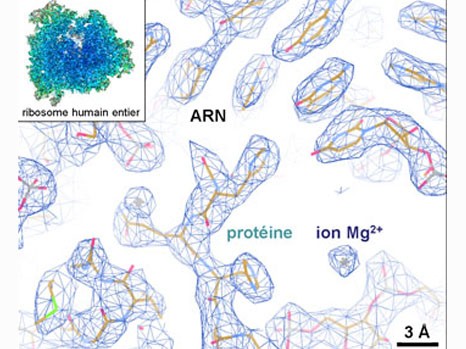
The study, published in Nature Genetics, is aimed at characterising malignant tumours that are suspected of belonging to the sarcoma group but have remained unclassifiable until now. Molecular investigations have shown alterations in the SMARCA4 gene, which encodes one of the subunits of BAF complexes. These complexes are involved in regulating the structure of chromatin, a compacted form of DNA in the cell nucleus.
To obtain these results, two complementary and independent approaches were employed (see appended schema on page 4). The first, based on pathological analyses, was used by the researchers from the Cancer Research Center of Lyon (Inserm/CNRS/Lyon 1 University/Léon Bérard Centre), and focused on unclassified sarcomas based on their microscopic features.
In parallel, the researchers from Institut Curie undertook a process of “blind” characterisation, analysing expression profiles by carrying out high-throughput RNA sequencing on approximately thirty undifferentiated sarcomas. Franck Tirode, an Inserm Research Fellow and codirector of the study, thus identified in this cohort a subgroup of tumours with very similar expression profiles, suggesting a common nature.
The Paris and Lyon researchers then pooled their results, having observed that they were interested in the same type of pathology. They validated these promising preliminary results by finding other similar cases or observations within one year, thanks to the collaboration of many hospital facilities distributed throughout France.
A new tumour entity
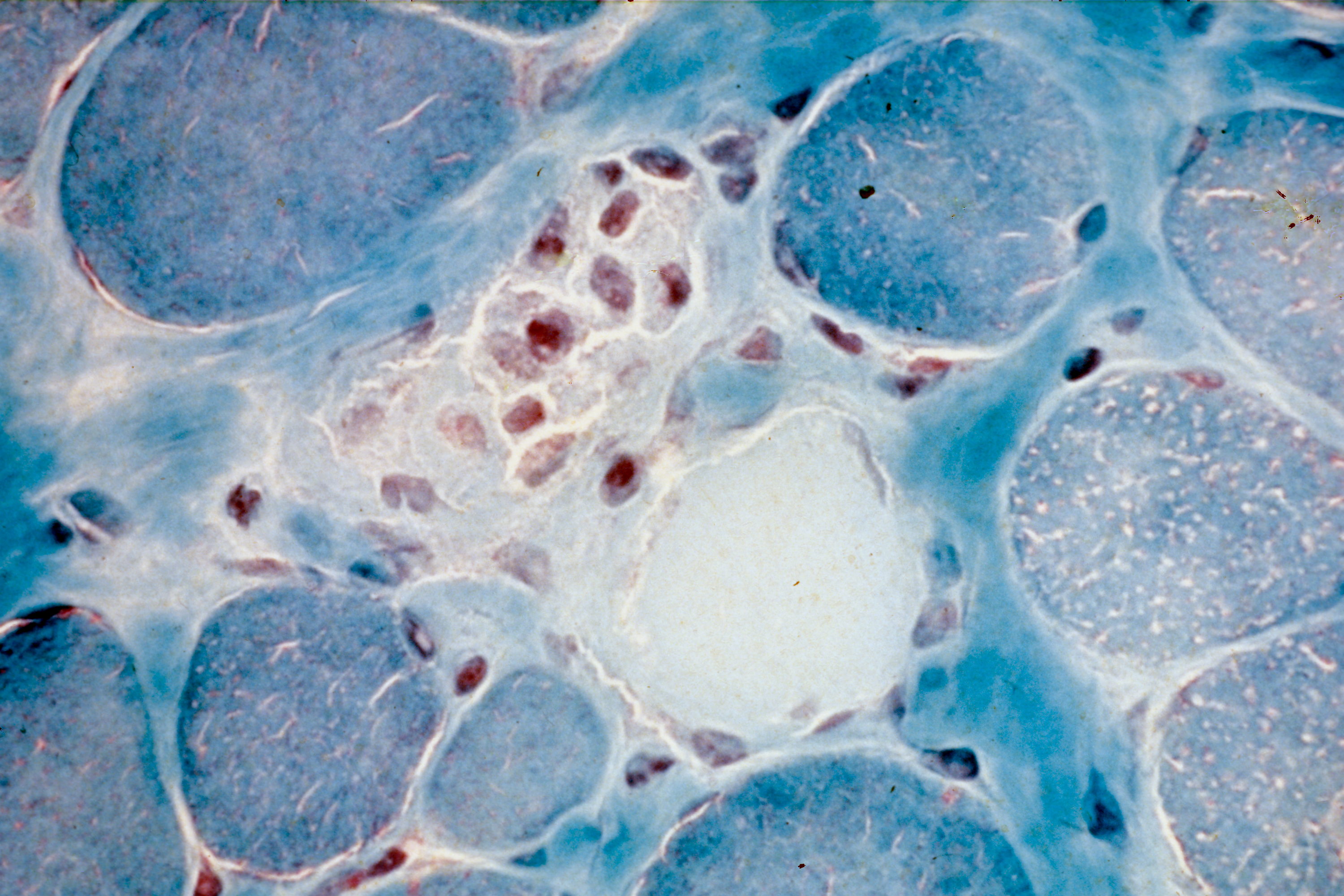
This collaborative study ultimately made it possible to identify 19 tumour samples with inactivation of the SMARCA4 gene. These tumours showed similar clinical and pathological features, producing large tumour masses that compressed the airways and progressed very rapidly, usually in young male smokers. Furthermore, when viewed under the microscope, they were all very similar to rhabdoid tumours, a form of paediatric cancer with a very poor prognosis. Moreover, these same tumours, described nearly 17 years ago by Olivier Delattre’s team at Institut Curie are, remarkably, associated with mutations of the SMARCB1 gene, which encodes a subunit of the BAF complex, of which SMARCA4 is a component.
The researchers compared expression profiles of all their tumours with numerous other types of tumours, and confirmed not only the exceptional homogeneity of these tumours among themselves, but especially the relationship of their signature to rhabdoid tumours. “However, although the oncogenic event might be comparable, the genomic complexity of the tumours we have studied differs considerably from the simple genomics of rhabdoid tumours,” says Franck Tirode.
Based on these results, the scientists could thus confirm that their cohort corresponded to a new homogeneous entity, which they have named “SMARCA4-deficient thoracic sarcoma.”
Better diagnosis for faster management
The researchers emphasise that these tumours are highly aggressive and resistant to current treatment methods. The BAF complex shows alterations in nearly 20% of human cancers, and is the subject of intensive research as a potential therapeutic target. Although this publication does not provide an offer of treatment to affected patients in the immediate future, it does show that these tumours are easily recognisable in clinical practice. Indeed, it was possible to identify new “prospective” cases during the study, enabling speedier management of the patients.
It was for this purpose that the authors of the study validated the SOX2 biomarker, which is particularly overexpressed by this new tumour entity, thus facilitating clinical diagnosis. They emphasise that the aggressiveness of these tumours is probably not unconnected with the overexpression of SOX2, which can endow cells with stem cell properties.
In conclusion, this work is part of the present concept of oncology and “personalised therapy,” in which the establishment of an accurate diagnosis is an essential prerequisite for the success of precision medicine.
SarcomasSarcomas are rare tumours that represent fewer than 1% of all new cancer cases, i.e. approximately 3,500 new cases a year. They affect people of all ages, although they are more frequently seen in children and young adults. They are found in bones, cartilage, adipose tissues, muscles, blood vessels or other connective or supporting tissue. Diagnosis of these rare tumours is coordinated at national level by the Groupe Sarcome Français (French Sarcoma Group) network, comprising 3 regional referral centres, including the Bergonié Institute (Bordeaux), Léon Bérard Centre (Lyon) and the Gustave Roussy Institute (Villejuif).
[1] Cancer Research Center of Lyon, Inserm U1052/CNRS 5286, Claude Bernard University Lyon 1, Léon Bérard Centre, “Therapeutic Targeting of the Tumor and of its Immune Environment” team
[2] Institut Curie, Inserm U830, “Genetics and Biology of Paediatric Cancers” team






 Human embryonic stem cells have the potential to form in vitro neural tube -like structures of the embryo. ©Inserm/Benchoua Alexandra
Human embryonic stem cells have the potential to form in vitro neural tube -like structures of the embryo. ©Inserm/Benchoua Alexandra