A team of researchers from Inserm led by Fanny Mochel and located at the Brain and Spinal Cord Institute (Inserm/CNRS/UPMC/AP-HP) has just demonstrated the therapeutic potential of triheptanoin in ten patients with Huntington’s disease. Derivatives of this triglyceride, with its unique composition, might be able to slow the progression of the disease by improving the energy metabolism of the brain. This research is published in the journal Neurology.
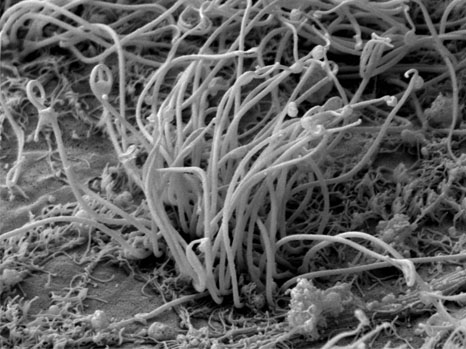
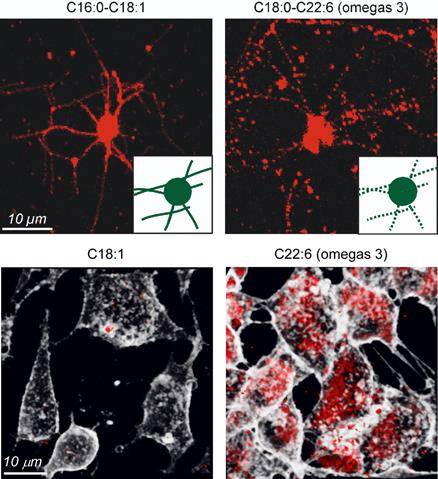
Study of the Huntington’s disease. © Inserm/Saudou, Frédéric
Huntington’s disease is a genetic disease that affects approximately 5,000 people in France, with another 10,000 individuals at risk of developing it. Mutation of the gene encoding the huntingtin protein results in a progressive degeneration of the neurons, especially in regions of the brain involved in the control of movement, thereby causing serious neurological, motor, cognitive and psychiatric problems. Weight loss is also observed in patients at an early stage in the disease, despite normal or even increased food intake. These two observations (degeneration of neurons and loss of weight) led the researchers to propose the hypothesis that an energy deficit in these patients might play an important role in the onset and progression of the disease symptoms.In order to validate this hypothesis, the researchers had to succeed in measuring energy fluctuations in the brain during certain brain activation states. To do this, they needed to develop technologies enabling the measurement of compounds that reflect the energy metabolism of the brain in a dynamic manner.
“We observed that variations in the relative concentrations of two phosphorus-rich energy compounds (Pi/PCr) perfectly reflect this dysfunction in brain energetics,” explains Fanny Mochel, who coordinated this work.
In practical terms, this relationship was measured using NMR.[1] While energy production in their brains was being recorded, the nine patients and thirteen healthy volunteers were asked to perform a simple mental “effort” task causing their brains to consume energy. For this, they were subjected to a visual stimulus in the form of a flashing checkerboard for several minutes.
The Pi/PCr ratio was clearly reduced in patients compared with control subjects, indicating that their brain metabolism was impaired when faced with this effort. Once this biomarker was validated, the researchers were then able to test the efficacy of potential treatments in re-establishing normal brain metabolism.
A pilot clinical trial…
For one month, ten patients with Huntington’s disease underwent a treatment based on triheptanoin, a triglyceride (in the form of an oil to be taken with meals), in order to improve energy metabolism in their brains. At the end of this therapeutic trial, their energy profiles had become normal. This treatment was not chosen at random, since it had already proved effective in patients with rare diseases who were unable to use certain lipids present in their diet. The idea was to compensate for generalised energy dysfunction by supplementing the diet with specific metabolites that could restore the body’s energy metabolism to normal. The novelty of the approach proposed by Dr Mochel is the application of this energy treatment concept to diseases that affect the brain, such as Huntington’s disease.
…Prior to a larger-scale trial.
On the basis of these results, the researchers will launch a one-year double-blind trial in France and the Netherlands at the beginning of 2015, in approximately one hundred patients, using clinical and imaging parameters as assessment criteria.
“The complexity of the disease obliges us to design additional therapeutic approaches in order to target different aspects—genetic, metabolic, inflammatory, etc.—of the disease. Used together, these approaches will probably have the best chances of success, such as happens with other, more common diseases,” concludes Fanny Mochel.
These results, which are patent-protected, were the subject of a recent licensing agreement negotiated and signed by ICM (the Brain and Spinal Cord Institute) and Inserm Transfert with the American company Ultragenyx, which will launch a new clinical trial in the area of Huntington’s disease.
[1] Nuclear magnetic resonance (NMR) is a noninvasive method for studying the biochemistry and metabolism of the central nervous system. It enables the accurate quantification of several tens of molecules. (Source: French Atomic Energy Commission, CEA)